RIC Goal
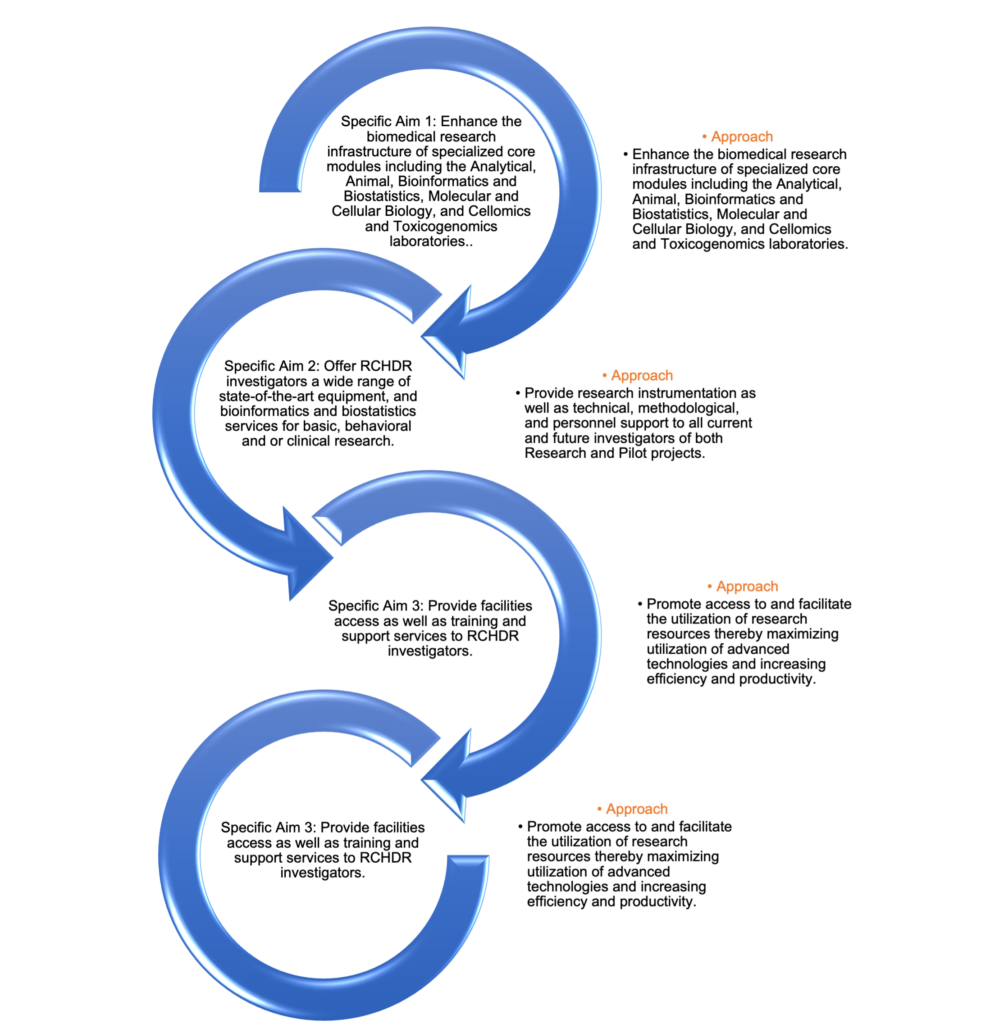
Ø The RIC’s overarching goal is to provide a robust infrastructure, research resources, and scientific expertise to assist RCHDR investigators to implement innovative research aimed at improving minority health and reducing health disparities.
Ø The RIC at Jackson State University (JSU) RCMI Center for Health Disparities Research (RCHDR) is designed to accelerate discoveries in basic, behavioral, clinical, and/or translational research.
Ø The RCHDR RIC serves as a strong catalyst for basic biomedical, behavioral, and/or clinical research at JSU on important health disparity issues of concern to minorities and underserved communities.
Ø The RCHDR RIC provides critical support to increase the quality, productivity, cost-effectiveness, and biomedical research outcomes of all RCHDR investigators.

Key Personnel

RIC Director Jacqueline Stevens, Ph.D.
Co-Lead, Molecular and Cellular Biology Core
jacqueline.j.stevens@jsums.edu
601-979-3462

Yiming Liu, Ph.D.
Co-Lead, Analytical Core
yiming.liu@jsums.edu
601-979-3491

Ibrahim Farah, Ph.D.
Co-Lead, Animal Core Facility
ibrahim.farah@jsums.edu
601-979-3466

Hafiz A. Ahmad, Ph.D.
Co-Lead, Biostatistics Core
hafiz.a.ahmad@jsums.edu
601-979-4048

Hung-Chung “Joe” Huang, Ph.D.
Co-Lead, Bioinformatics Core
hung-chung.huang@jsums.edu
601-979-2189

Paul B. Tchounwou, Sc.D.
Co-Lead, Cellomics & Toxicogenomics Core
paul.b.tchounwou@jsums.edu
601-979-0777
Essential Functions/
Services Research Resources
Major Scientific Equipment and Technologies
Tissue/Cell Culture
Two dedicated tissue culture rooms are available. Cell culture equipment include: 3 biosafety cabinets with UV light, 2 cryosafe liquid nitrogen tanks; 2 -80oC deep freezers, 2 4oC refrigerators, 2 refrigerated microcentrifuges, 2 humidified CO2 incubators, 2 cell disruptors, and 2 Olympus inverted light microscopes equipped with Dell workstations, CellSens software, and digital cameras. Additionally, a walk-in cold room (-4oC) is available on site. Two large capacity autoclaves (Primus Model PSS5-A-MESD) are in place to sterilize all materials or waste that need to be treated before use or disposal.
Biomedical Imaging
Olympus FV1000 confocal laser scanning microscope system with an inverted IX81 microscope featuring improved basic performances (sensor system, scanning system, and illumination system performances). This microscope is equipped with three fluorescence channels, three lasers, and AOTF to meet various applications in a wide range of advanced research fields.
Leica TCS SP2 Confocal microscopy system that uses high-intensity laser light and a confocal aperture to obtain optically focused images from fluorescent samples.
JEOL-1011 Transmission Electron Microscope system equipped with is 0.2 nm lattice resolution with a magnification of 50 to 1,000,000 under the accelerating voltage of 40 to 100 kV, a camera system, and a computer with LC monitor.
Quanta 200 Environmental Scanning Electron Microscope (ESEM) equipped with a CCD IR camera and a Pentium PC with LCD monitor.
Tuscan LYRA3 Focused Ion Beam Scanning Electron Microscopy system (SEM/FIB) equipped with 2 closed-loop nano-manipulators and an electron dispersive X-Ray spectroscopy analyzer (EDX) for elemental analysis.
Olympus Epifluorescence microscopy system equipped with Comet assay software for DNA damage analysis.
Bio-analytical Analysis and other Functional Assays
Available equipment include: Horiba FluoroMax4 fluorescence spectrofluorometer, Perkin Elmer FT-IR spectrometer, Shimadzu UV-2600 UV-Vis spectrophotometer, Shimadzu Model UV-3700 Solid Spec UV-VIS-NIR spectrophotometer, Shimadzu Model AA-6701 atomic absorption spectrometer, Shimadzu Prominence HPLC with auto sampler and UV and fluorescence detectors, Agilent 7100 Capillary electrophoresis with UV detection, Thermo Model TSQ Quantum GC-MS/MS /LC-MS/MS, Varian Model 820-MS LC-ICP-MS, Shimadzu GC/ECD and FID detectors, Zeta sizer model 3600, Costech Model 4010 elemental combustion system/analyzer, Tri-Carb 4910TC liquid scintillation counter, and Brucker 500 MHz NMR spectroscopy system with analytical probes.
Flow Cytometry Analysis
There are two major pieces of flow cytometry equipment available for cell sorting and functional analysis of phenotypes and contents of eukaryotic cells. These include the Becton Dickinson Biosciences FACS Calibur and FACS Lyric fluorescence activated cell sorting systems that provide services such as cell cycle analysis, apoptosis assessment, and cell protein detection. In addition, we have a Cellometer Vision analyzer that allows us to perform analysis of cell cycle distribution.
Gene Expression Analysis
The RCHDR RIC hosts the following equipment: Illumina RNASeq 550 System, automatic environmental speed Vac system, BioRad molecular image gel documentation system, Eppendorp mastercycler gradient thermocycler, PCR Workstation, Stratagene Stratalinker 1800 UV crosslinker, BioRad gel dryer, BioRad UV spectrophotometer, Precision incubator/oven, Omni tissue homogenizer, electrophoresis units, Amersham electro-blotting system, Olympus inverted phase contrast microscope with camera, Affymetrix Microarray system, Olympus epifluorescence microscope with camera, BioRad microplate reader, Labconco™ FreeZone™ Freeze Dry System, and Beckman ultracentrifuge.
In-vivo Experimentation with Laboratory Animals
The research resources available in the Animal Core Facility include two animal husbandry rooms (1 for rats and a for mice), large-scale cage washer, large size freezer, humidity and temperature control equipment, various regular racks and cages for rats and mice, special metabolic cages and racks, a ventilation system, both constant and portable air filtration systems, light-dark cycle controls, anesthesia, surgery, post mortem, euthanasia and sampling station; manual food and water dispensers, food and bedding storage and a general storage area, enviro-cages for immunodeficient animals, laminar flow hood, and autoclave.
Bio-statistical Support in Study Design, Data Management and Data Analysis
The Biostatistics unit of RCHDR provides comprehensive analytical services, from designing experiments to data analysis using current software mainly through one-to-one consultations and workshops. The unit has adopted new procedures and methodologies in latest Big Data trends using artificial intelligence and related approaches to model disease propagation and health disparities. This unit provides services and workshops in developing innovative research and pilot project proposals, particularly using new modeling approaches, such as neural network and/or genetic algorithm in addressing new trends in data analysis. The Biostatistics lab is currently equipped with 25 Dell OptiPlex desktop computers with Intel core i7 processors running on Windows 10 operating system. All computers carries latest versions of statistical and modelling software, such as SAS, SPSS, SysStat, Minitab, Neuroshell, @Risk, and Matlab. The lab has a JTouch 65 inch whiteboard with capacitive touch and overhead projectors for training and presentations.
Bio-informatics and Computational Biology support

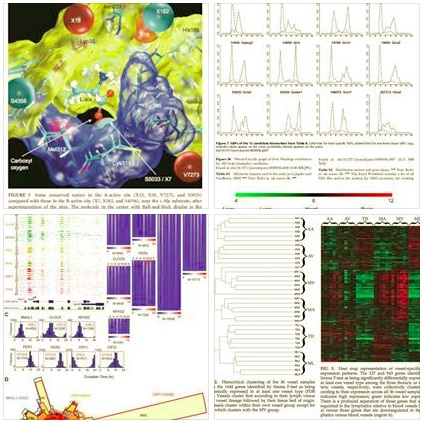




The Bioinformatics and Computational Biology support unit offers regular training workshops and services to RCHDR investigators and students. Training workshops focused on bioinformatics databases, molecular modeling and simulations, next generation sequencing (NGS) and microarray datasets, genome-wide SNP and eQTL analyses, and biomarkers discovery for disease diagnosis and prognosis, to cover both genomic and structural bioinformatics research. An emphasis is put on the understanding of the relationships between genome and phenome (e.g., transcriptomics, proteomics, and metabolomics phenome data). Most NGS data analyses (e.g., on ChIP-seq and RNA-seq data) are supported. The Bioinformatics unit is equipped with 10 Dell Precision Tower 7910 workstations with up to 64GB RAM, and a Linux server (Ubuntu 18.04LTS) containing 128GB RAM and GPUs with more than 500 streaming processors for supercomputing and bioinformatics research. Recently we have established a Big Data graduate program (Computational and Data-enabled Science and Engineering PhD program), and acquired an account with the Extreme Science and Engineering Discovery Environment (XSEDE) supercomputing resource (https://www.xsede.org). This account provides remote access to 68672 CPUs and 2.5 TBs of temporary data storage on XSEDE site, and allows for 100,000 hours of supercomputing time. Aligning these computational resources allows the RCHDR RIC to accommodate the needs of researchers within various disciplines, and facilitate the cost-effective implementation of big data-related projects.